|
It also means that toggling different targets of interest beyond the conventional SELEX procedure can isolate cross-reactive aptamers with broad range of binding 30, 31. coli, implicating that some bacterial strains share common targets on their cell surface 29. coli strain demonstrated that the selected aptamers can also bind other strains of E. A previous study which performed SELEX against a single E. The dissociation constants ( K d) of several reported aptamers are in the nanomolar range with low limit of detection of about 1 to 1000 CFU/mL 26.Īlthough most bacterial cell-SELEX studies focus on targeting single bacterial strain, it is possible that aptamers could bind epitopes commonly shared among different bacterial strains 27, 28. Using bacterial cell-SELEX, several aptamers have been isolated against different bacterial species with high affinity and specificity. Using cell-SELEX, aptamers can be isolated against target epitopes in its natural physiological context, even when the exact target on the cell surface is unknown 24, 25. In this study, nucleic acid aptamers are proposed for the development of inexpensive and rapid diagnostic tools, because aptamers could recognize target epitopes with high selectivity and specificity using the screening technique of systemic evolution of ligands by exponential enrichment (SELEX) 23. This suggests that the detection of bacterial OMVs is more effective than the detection of bacterial cells in clinical blood samples 22. Detecting living bacteria in blood is difficult due to the immune regulation of host cells and the serum bactericidal activity 20, 21. Despite the vast amount of research, sensitive bacterial detection methods which meet commercial demands in real situations are yet to be developed 19. Conventional methods of bacterial detections in general rely upon three types of laboratory-based techniques such as optical assays using fluorescent labeling 13, 14 or surface plasmon resonance 15, mechanical quartz crystal microbalance sensors 16, 17 and electrochemical sensors 18 such as potentiometric, amperometric and impedimetric assays. These antigenic moieties, including LPS 8, 9, peptidoglycans 10, 11 and lectins 12 have served as substrates of the bioreceptors. Various surface antigens exist on the cell surface of Gram-negative bacteria, which can be potential targets for bacterial detection. OMVs are known to trigger severe pathogenesis, enhance bacterial survival, transfer genetic and protein components for cell-free intercellular communication, deliver toxic compounds, and trigger immune response in host cells 6, 7. Furthermore, outer membrane vesicles (OMVs) with size of 20 to 250 nm, which are commonly produced and secreted from outer membranes of bacteria, carry various bacterial membrane proteins and other virulence factors 4, 5. Lipopolysaccharides (LPS), a major constituent of the outer membrane of Gram-negative bacteria, induces a strong innate immune response in host environments 3. 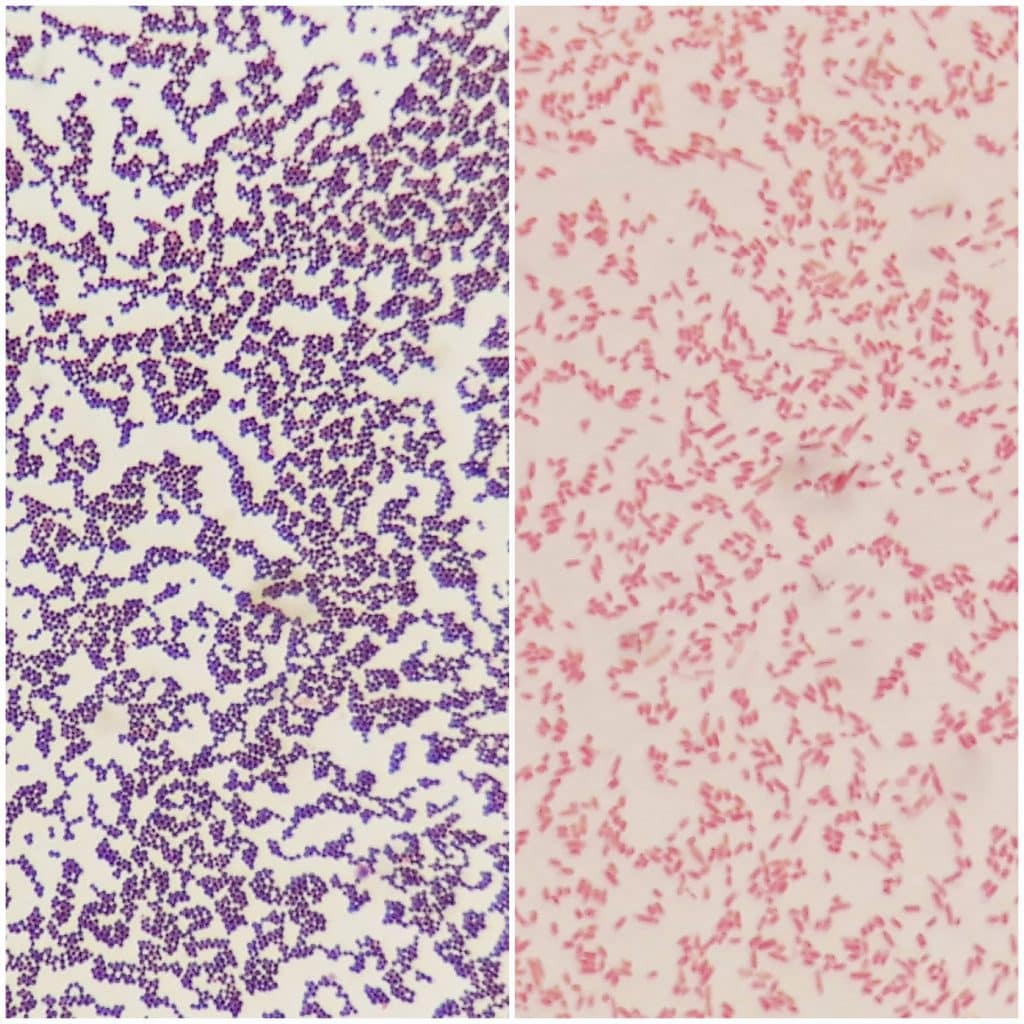
They are resistant to various antibiotics because their outer membrane evolves and adapts to the protective mechanisms against the selection pressure of antibiotics 2. Gram-negative bacteria especially represent an important medical challenge. Aptamers developed in this study show a great potential to facilitate medical diagnosis and early detection of bacterial terrorism, based on the ability to detect bacterial OMVs of multiple Gram-negative bacteria.īacterial infections can be detrimental due to their virulence factors, underscoring the need for development of rapid, accurate, and sensitive detection techniques 1. 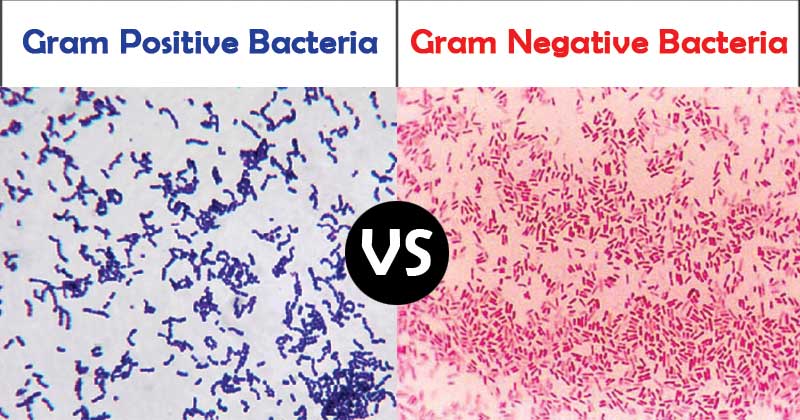
The GN6-ELAA showed high sensitivity to detect as low as 25 ng/mL bacterial OMVs. Using GN6 aptamer, we constructed an Enzyme-linked aptamer assay (ELAA) to detect bacterial outer membrane vesicles (OMVs) of Gram-negative bacteria, which contain several outer membrane proteins with potent immunostimulatory effects. Two aptamers selected, GN6 and GN12, were 4.2-times and 3.6-times higher binding to 10 8 cells of Gram-negative bacteria than to Gram-positive bacteria tested, respectively. coli K12, and Serratia marcescens, were sequentially incubated with the pool of random DNA sequences at each SELEX loop. Three Gram-negative bacteria, Escherichia coli DH5α, E. In this study, we isolated single-stranded DNA aptamers against multiple Gram-negative bacterial species using Toggle-cell-SELEX (systemic evolution of ligands by exponential enrichment) and constructed an aptamer-based detection tool towards bacterial secretory cargo released from outer membranes of Gram-negative bacteria. Despite the research advances, current identification tools still exhibit limitations in detecting Gram-negative bacteria with high accuracy. Infection of various pathogenic bacteria causes severe illness to human beings.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
